Fully automated template matching method for ECG-free heartbeat detection in cardiomechanical signals of healthy and pathological subjects
Executive Summary
This study introduces a fully automated template matching method for ECG-free heartbeat detection in cardiomechanical signals, validated on 256 SCG, GCG, and FCG signals from 150 subjects, including 95 pathological cases. The method employs a novel automatic template selection algorithm, achieving sensitivity and PPV of up to 99.2% and 99.3% for healthy subjects and 85% and 95% for pathological subjects. Statistical analyses demonstrated high accuracy in inter-beat interval estimation, with limits of agreement within ±6 ms for healthy subjects and ±13 ms for pathological subjects, surpassing all prior ECG-free methods in performance and reproducibility.
Answer Machine Insights
Q: What is the main advantage of the proposed method over previous ECG-free heartbeat detection methods?
The method automates template selection, eliminating the need for manual intervention and ensuring reproducibility.
The novel automatic template selection algorithm was specifically designed to overcome the limitation of manual template selection, thus ensuring reliable and reproducible results without requiring the intervention of a skilled operator.
Q: How does the method perform on pathological subjects?
It achieves sensitivity and PPV of 85% and 95%, respectively, despite the challenges posed by irregular signal morphologies.
On pathological subjects, it scored a sensitivity of 85% and a PPV of 95% for both SCG and GCG.
Key Results
Sensitivity and PPV of 97.8% and 98.6% for SCG signals in healthy subjects.
Limits of agreement for inter-beat intervals within ±6 ms for healthy subjects and ±13 ms for pathological subjects.
Visual Evidence
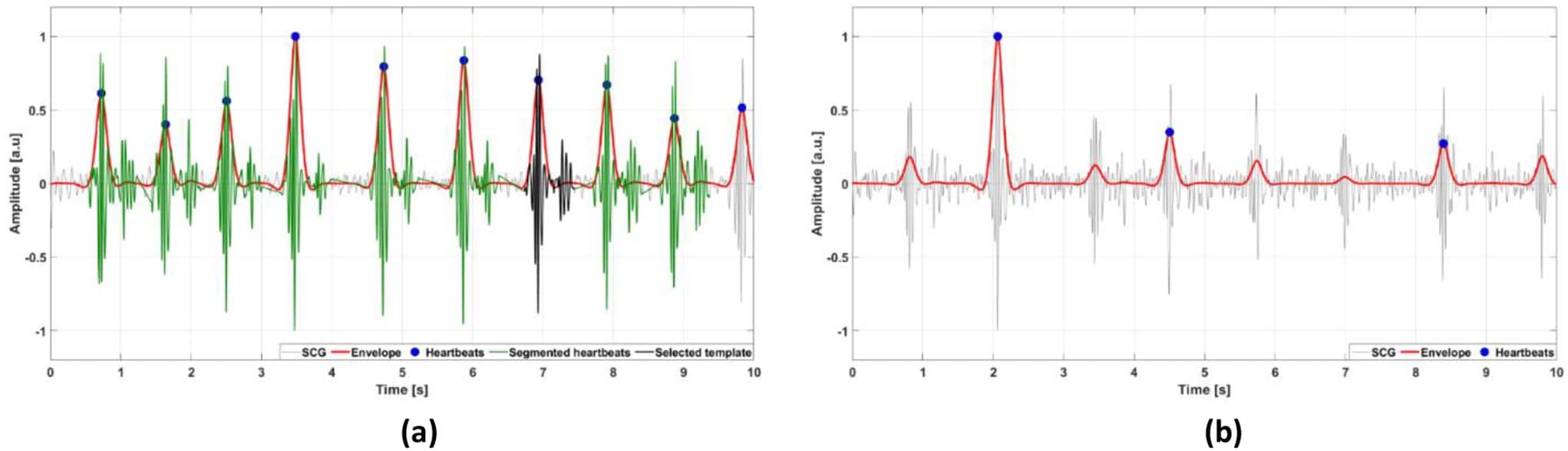
Fig. 3 Examples of 10-second segments of SCG signals classified by the automatic template selection as: (a) reliable; (b) non-reliable. The SCG signals are depicted as thin grey lines, the envelope signals as thick red lines, the locations of heartbeats (envelope peaks) are marked with full blue circles, the segmented heartbeats are depicted as thick
Clinical Snapshot
Evidence Rating
Relevance
high Priority